Ringach-Shapley-Hawken 2002
Macaque V1: Orientation Tuning Bandwidth and Circular Variance
| Model Name | Model Details | Benchmark Scores* | |||
| Bandwidth | Circular Variance | ||||
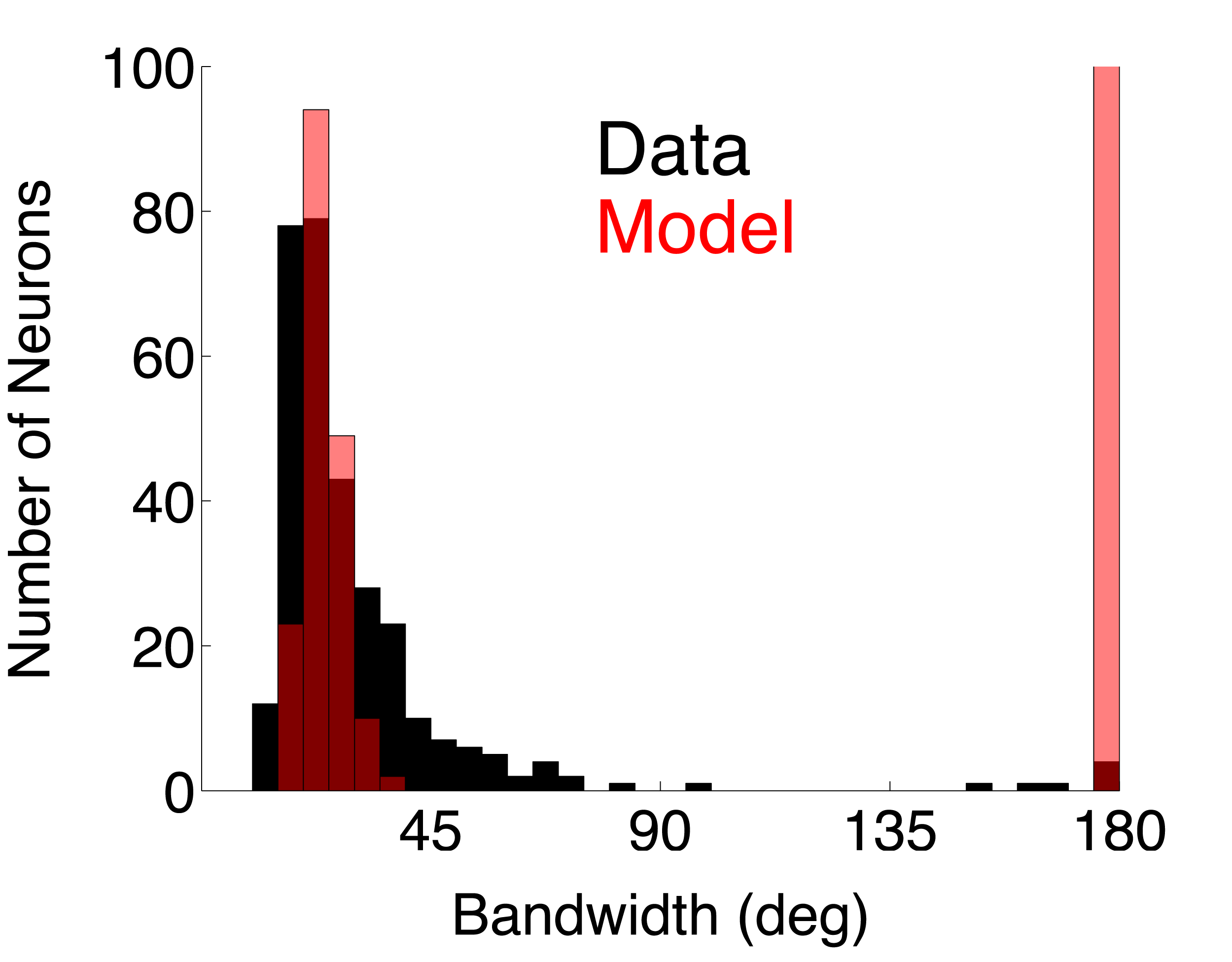
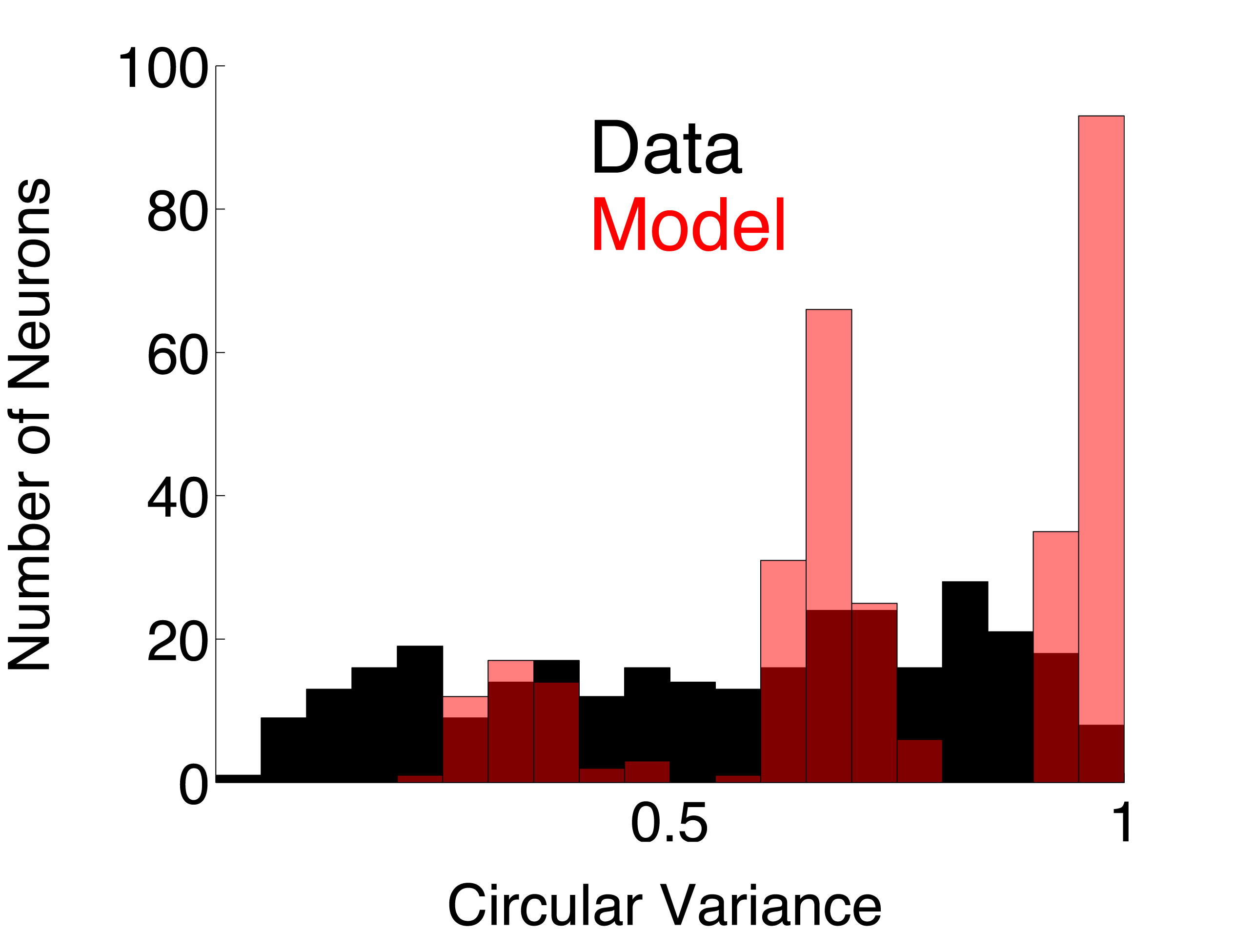
| DS_Post_Fac | Direction selective complex cell population using post-synaptic, facilitatory DS interaction (model page) | 0.4053 | 0.3761 | ||
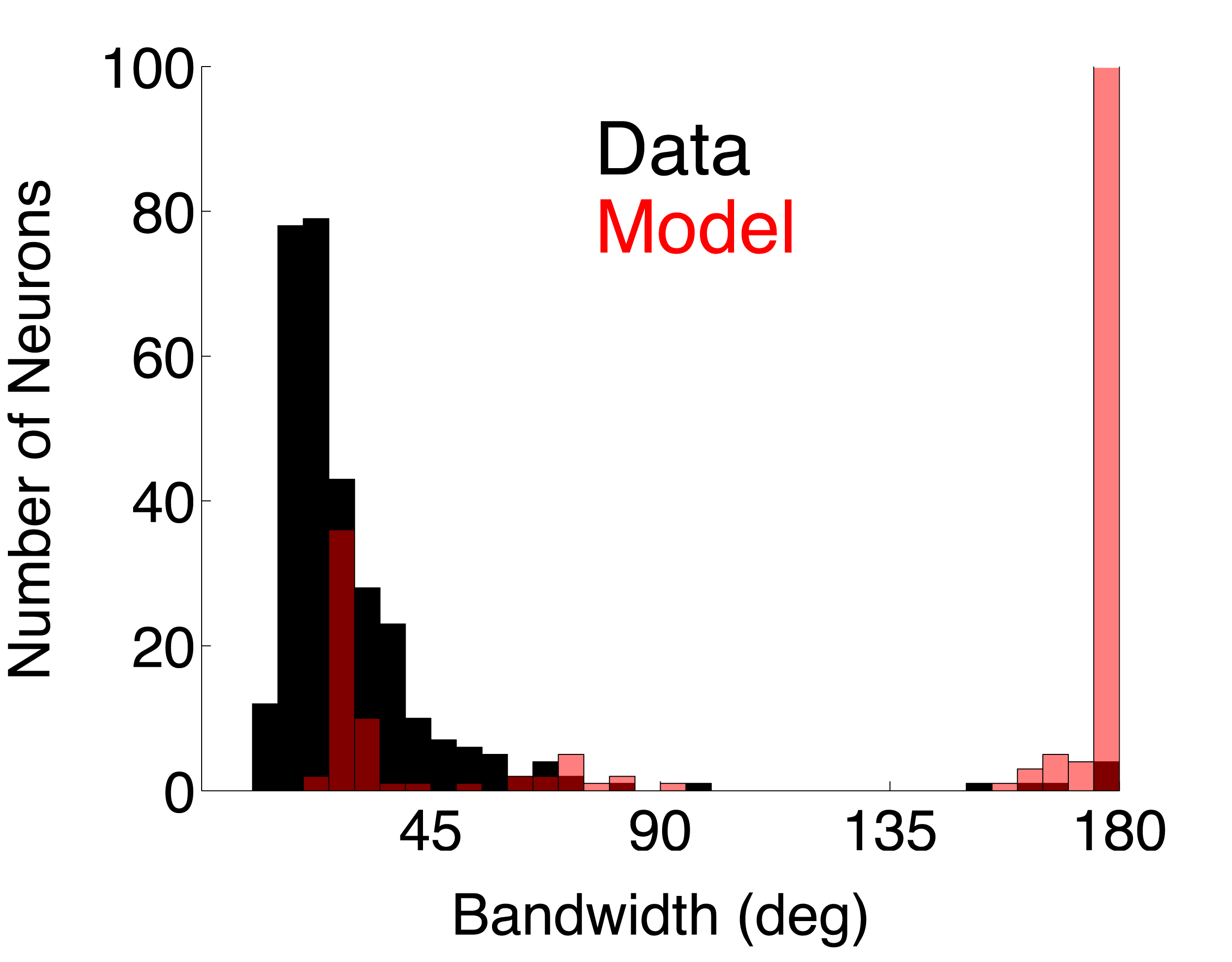
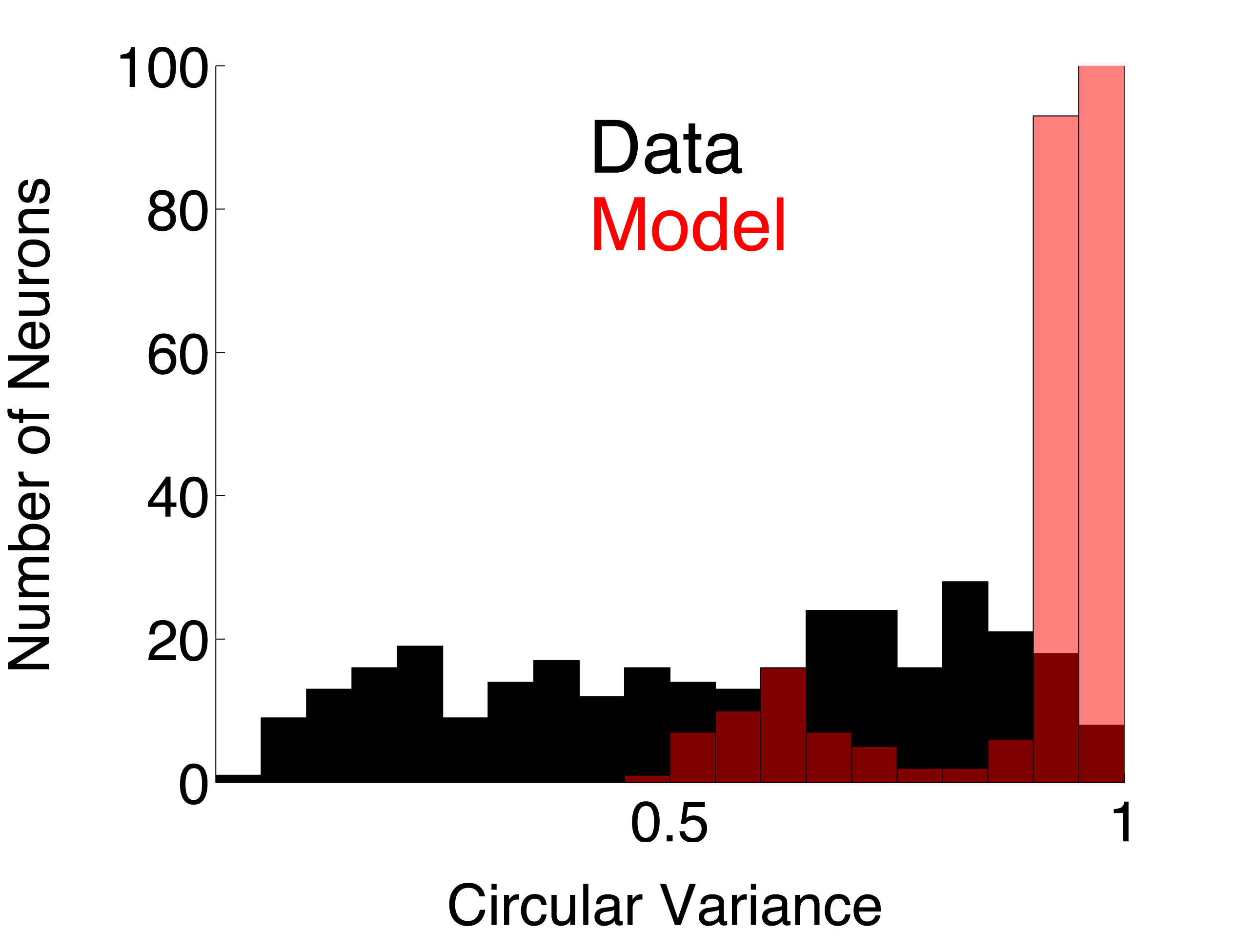
| DS_Simp | Direction selective simple cell population using linear summation ( model page) | 0.7828 | 0.7341 | ||
Stimulus Protocol
The stimulus protocol consists of drifting sinusoidal gratings optimized for SF, TF, size and contrast. Stimulus orientation was varied in 15 degree steps. Each trial lasted for 4 seconds. To view this stimulus, click the button below to launch the iModel stimulus viewer.
Description of the Sampled Population
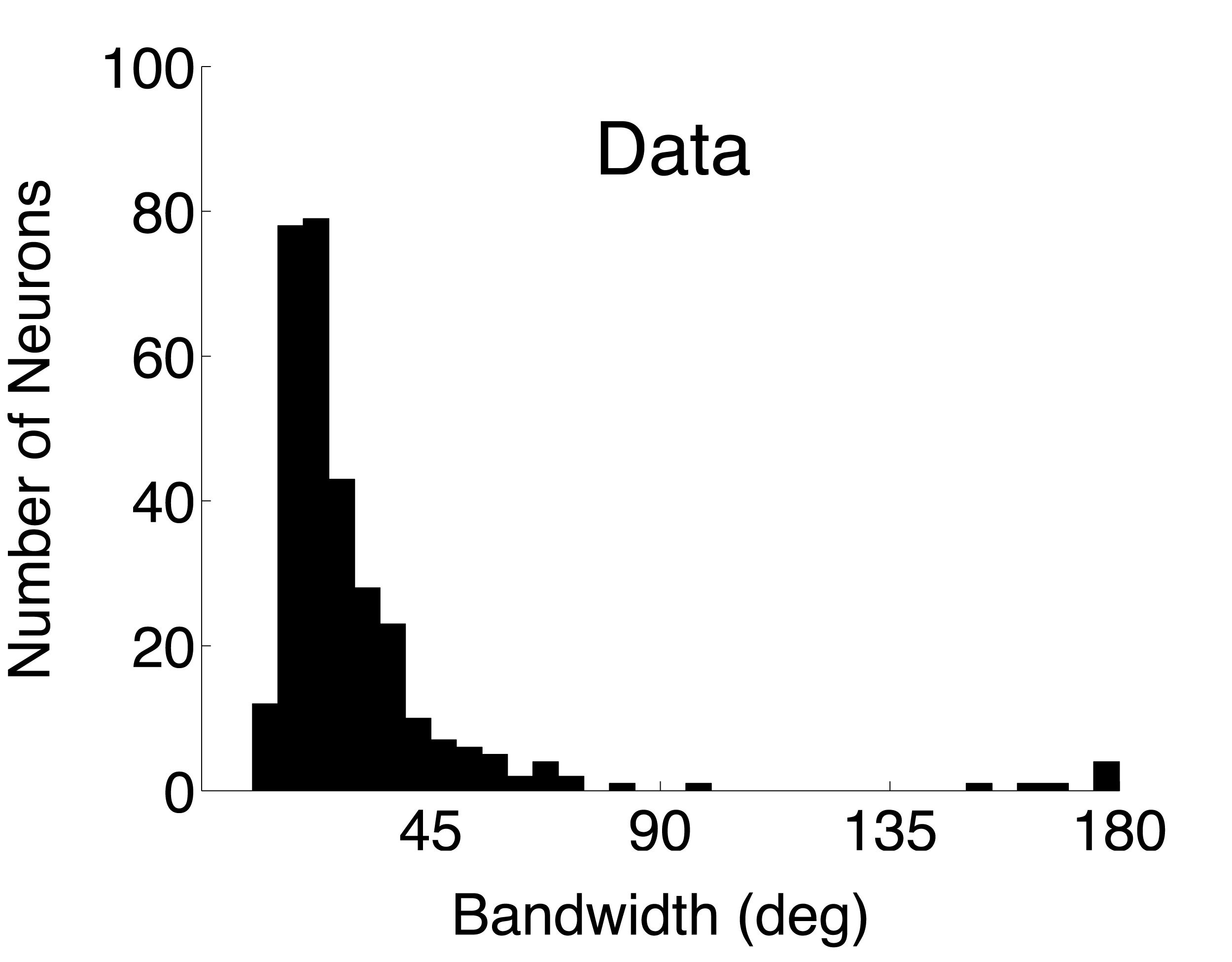
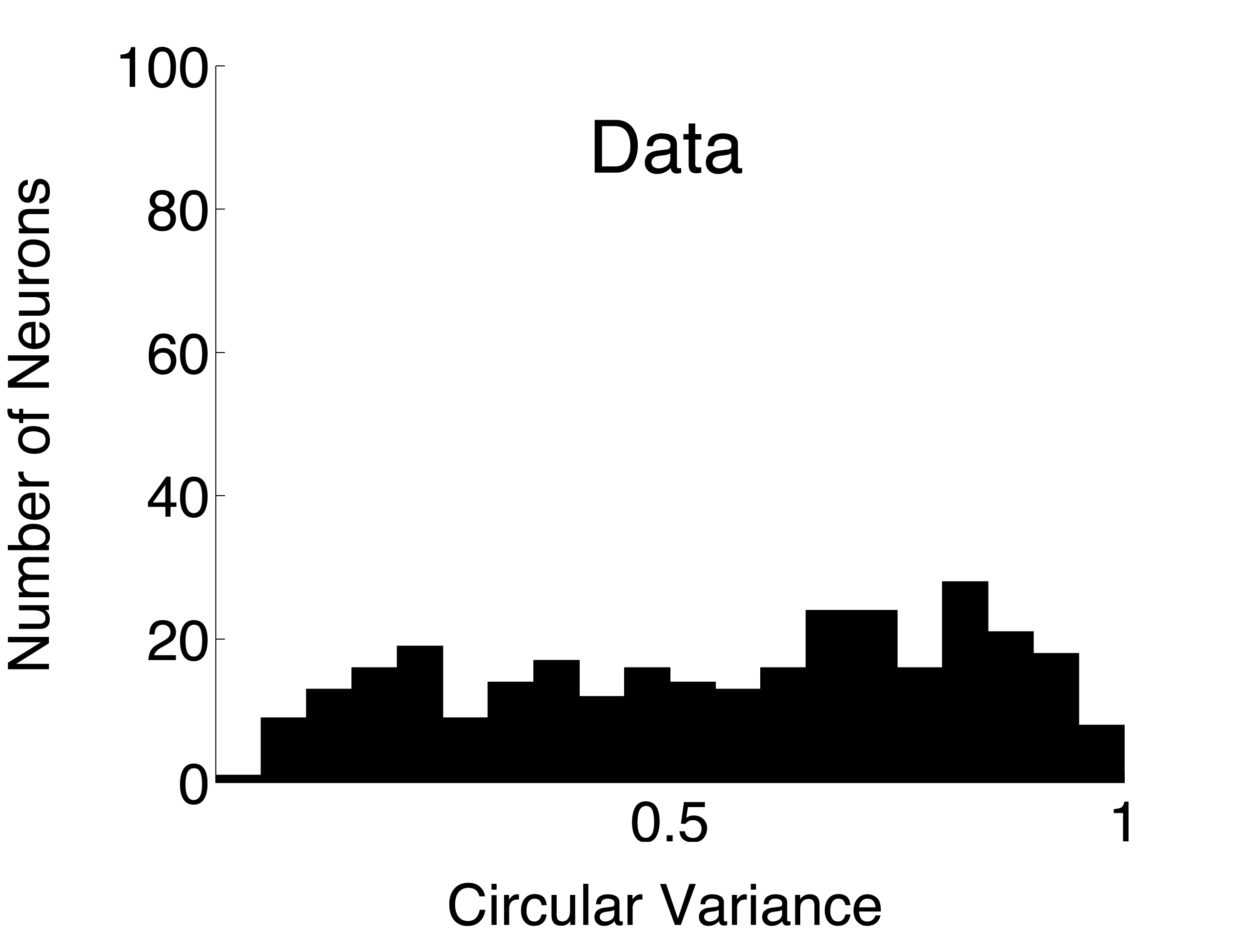
The paper presents bandwidth and circular variance data collected from 308 V1 neurons in 26 anesthetized macaques. Both simple and complex cells were lumped together in the analysis. Cells were distributed across all layers of cortex. The authors have made their data freely available at http://www.ringachlab.net/lab/Data.html.
Data Analyses
For each single unit, a direction tuning curve is computed that represents the mean firing rate (without baseline subtracted) at each direction (orientation) tested. Click on the analysis names to view the C source code for each analysis.Bandwidth. Orientation tuning bandwidth (BW) was computed from the direction tuning curve for each unit by first interpolating the tuning curve at a one degree resolution using a Hanning (cosine) window that had a half-width at half-height of 13.5 deg. The location of the peak of the interpolated tuning curve was determined, and the orientation angles closest to the peak for which the response fell to 1/sqrt(2) (or 70.7%) of the peak response on either side of the curve were estimated. BW was defined as one-half of the difference between these two angles. If the tuning curve never went below response criterion, the BW was set to 180 deg.
Circular Variance. The circular variance (CV) was calculated from orientation tuning curves as follows. Each point on the orientation tuning curve is considered to be a vector where the response amplitude is the vector length and the direction of motion is the vector angle. The vector, S, that is the sum of all of the tuning curve vectors is computed. The CV is the length of S divided by the sum of all the response amplitudes. Note that CV averages the responses for the two directions of motion at each orientation. If the tuning curve is flat (response = C for all directions), then CV = 1. If an orientation-tuned neuron is so selective that its response is non-zero for only one direction (or orientation) on the tuning curve, then CV = 0.
Benchmark Scoring
The score is computed as the Kolmogorov-Smirnov (K-S) test statistic for the equality of one-dimensional distributions. Lower numbers represent a better match between model and benchmark data.
References
- Ringach DL, Shapley RM and Hawken MJ (2002) Orientation Selectivity in Macaque V1: Diversity and Laminar Dependence. J Neurosci 22:5639--5651.
Note: This page is for demonstration only.
Step 1: Select benchmark to run
Step 2: Select modelRingach-Shapley-Hawken 2002 - Macaque V1: Orientation Tuning Bandwidth and Circular Variance
The benchmark defines the visual stimulus as well as other relevant parameters of the simulation (e.g. duration, number of trials).
Step 3: Select units in model to benchmarkHere would be a dropdown list for the user to select from publicly available models on the site, as well as user-defined models saved in their personal account.
Step 4: Run benchmarking simulationsHere would be a Response Selection applet for the user to select the model populations and/or sub-populations that they wish to benchmark against.
Once the stimulus, model and responses have been selected, the user will be directed to a login page for their iModel account. The simulation can then be sent to the cluster for execution. When finished, the user can access their account to view the data and run the Benchmark Analysis.